Structured Interfaces for Scientific Agents
Published:
TL;DR: this round is about scientific agents needing better interfaces to knowledge, tools, and intermediate workflow state. I picked four papers that make that interface explicit: a method-evolution graph for AI scientists, a framework that lets language agents call domain foundation models, a benchmark for hard literature discovery, and a controlled comparison of scientific-visualization agent interaction styles.
What I Am Watching This Round
The last few issues spent a lot of time on verifiable states, replayable workspaces, and hard evidence for data agents. I wanted this round to move one layer up. If agents are going to help with research, they need more than a browser, a search box, and a long context window. They need structured access to method lineage, specialist scientific models, full-text evidence, and workflow checkpoints that can be inspected before the final answer.
That is the common thread here. These papers are not saying that autonomous scientists are solved. They are mostly showing where the current interface is too thin: citation graphs do not explain method evolution, language-only agents mishandle non-linguistic scientific data, search agents miss paper-internal clues, and visualization agents either spend huge tokens on code or lose coherence in GUI workflows.
Paper Notes
Intern-Atlas: A Methodological Evolution Graph as Research Infrastructure for AI Scientists
Authors: Yujun Wu, Dongxu Zhang, Xinchen Li, Jinhang Xu, Yiling Duan, Yumou Liu, Jiabao Pan, Xuanhe Zhou, Jingxuan Wei, Siyuan Li, Jintao Chen, Conghui He, Cheng Tan.
Institutions: Shanghai Artificial Intelligence Laboratory; Peking University; Xi’an Jiaotong University; Zhejiang University; East China Normal University; Hunan University; Shanghai Jiao Tong University; Shanghai University; University of Chinese Academy of Sciences.
Date/Venue: April 30, 2026, arXiv preprint.
Links: arXiv | HTML
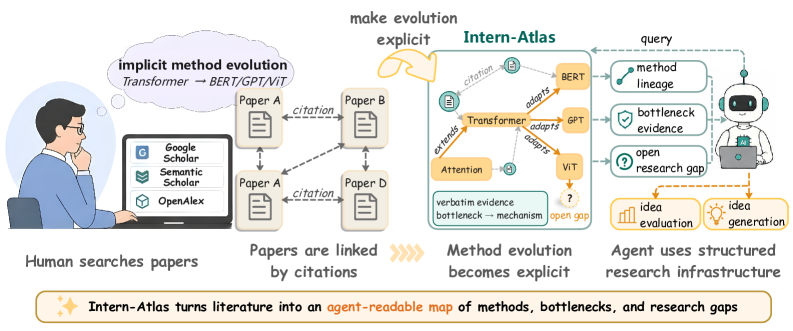
This overview is the best way into the paper. Intern-Atlas turns papers, methods, stubs, typed citation relations, and evidence snippets into a graph that an agent can query. The important claim is not that the graph replaces reading papers, but that method lineage becomes an object rather than something reconstructed privately inside a human researcher’s head. The caveat is that the graph still depends on LLM extraction, so the validator and benchmark matter as much as the graph scale.
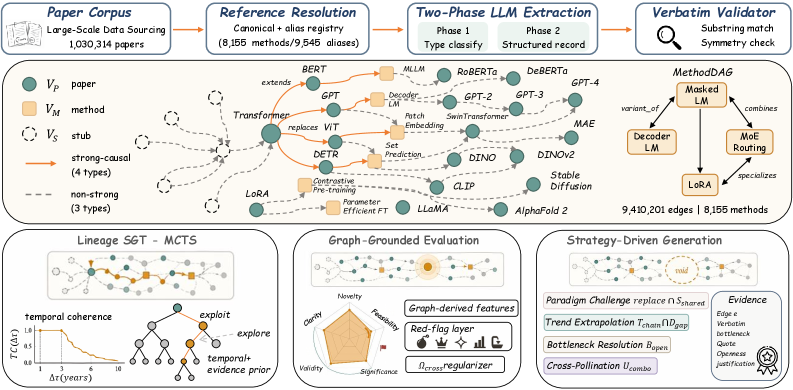
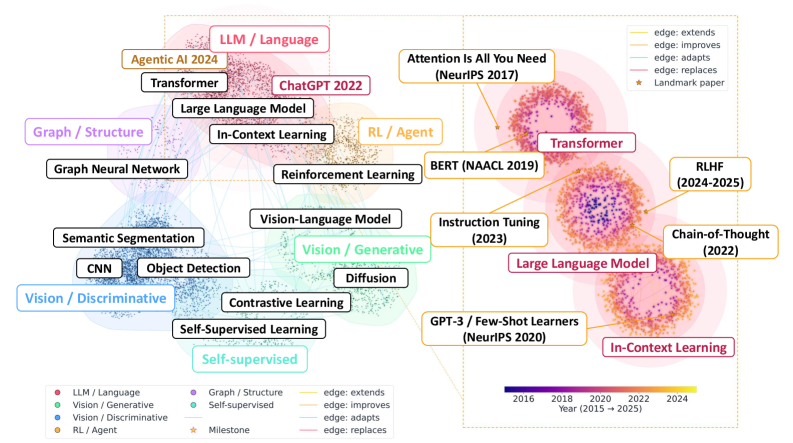
The landscape view shows why this is more than a citation graph. Methods cluster into broad research “continents”, and typed evolution edges connect landmarks such as Attention, BERT, GPT-3, chain-of-thought, instruction tuning, and RLHF. This figure supports the paper’s claim that agents need method-level topology, not only paper-level metadata. It should still be read as a representative visualization, since a readable map necessarily hides most nodes and edges.
The quality plot is where I start trusting the paper more. The authors compare against survey-derived method-evolution graphs and report 91.0% node match ratio, 89.7% edge reachable ratio, and 92.0% path semantic correctness. SGT-MCTS also recovers reference chains much better than beam search and random walks, with reported node recall, edge recall, and chain alignment all far ahead of the strongest beam baseline.
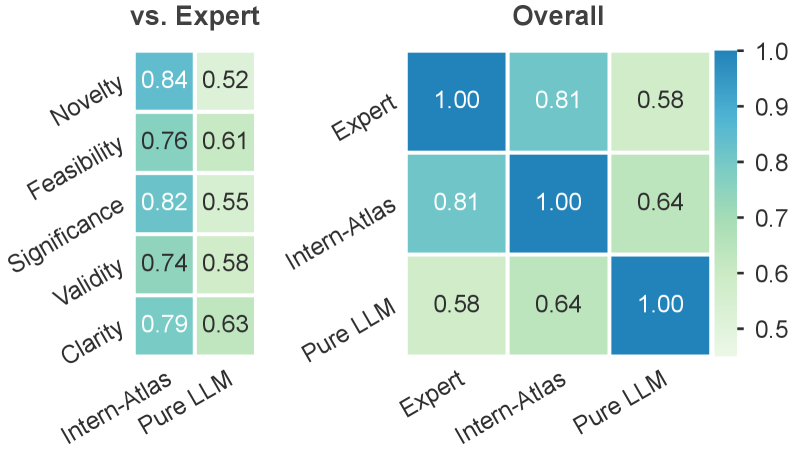
The downstream plot tests whether the graph helps with idea evaluation and idea generation. I like that the paper does not rely only on a pretty graph: it asks whether graph-grounded scores track publication strata and whether generated proposals win against baselines. The caution is familiar: idea quality is hard to evaluate automatically, so these results are evidence that the graph is useful, not proof that it can judge research taste on its own.
Quick idea: Intern-Atlas builds a method-evolution graph so AI research agents can query how methods extend, improve, adapt, or replace one another.
Why it matters: scientific search infrastructure is still mostly document-centric. Citation edges tell an agent that two papers are connected, but not whether one method solved a bottleneck in another. For research agents, that missing semantic layer is expensive. It forces the agent to infer method lineage from scattered prose, which is exactly where current systems hallucinate or overfit to familiar names.
Method walkthrough:
- The system starts from 1,030,314 AI publications from conferences, journals, and arXiv preprints, then resolves papers, canonical methods, aliases, and out-of-corpus stubs into graph nodes.
- It classifies citation-linked method relations into seven types: extends, improves, replaces, adapts, uses_component, compares, and background. The first four form the strong causal lineage graph.
- For causal edges, it extracts a bottleneck, mechanism, trade-off, and confidence score, with quoted spans checked by a deterministic substring validator.
- On top of the graph, it implements lineage reconstruction, graph-grounded idea evaluation, and gap-driven idea generation.
Evidence: the graph contains 8,155 canonical methods, 9,545 aliases, and 9,410,201 typed edges. The evaluation benchmark is built from 30 high-impact surveys and contains 2,268 nodes, 1,462 edges, and 133 evolution chains. The reported graph-quality scores are high, and the lineage search is much stronger with SGT-MCTS than with beam search. For idea evaluation, the overall score follows the expected order across extracted ideas from top-tier, core-conference, workshop, and rejected papers, with reported means of 8.48, 7.83, 6.85, and 5.84.
Why I care: this paper is close to Paper Radar’s own problem. A research assistant should remember more than a pile of PDFs and tags. It should know which methods were created to resolve which bottlenecks, and where an apparently new idea sits in the lineage.
Limitations/questions: I would not treat an LLM-built method graph as ground truth. The evidence spans help, but relation labels such as “improves” or “adapts” can still be context-sensitive. The next thing I would check is whether domain experts can edit the graph without breaking downstream operators, and whether smaller specialized fields show different error patterns from mainstream LLM/CV areas.
Connection to the tracked themes: data agents, research memory, automated scientific discovery, large-model-assisted research infrastructure.
Heterogeneous Scientific Foundation Model Collaboration
Authors: Zihao Li, Jiaru Zou, Feihao Fang, Xuying Ning, Mengting Ai, Tianxin Wei, Sirui Chen, Xiyuan Yang, Jingrui He.
Institutions: University of Illinois Urbana-Champaign.
Date/Venue: April 30, 2026, arXiv preprint.
Links: arXiv | HTML | project | code
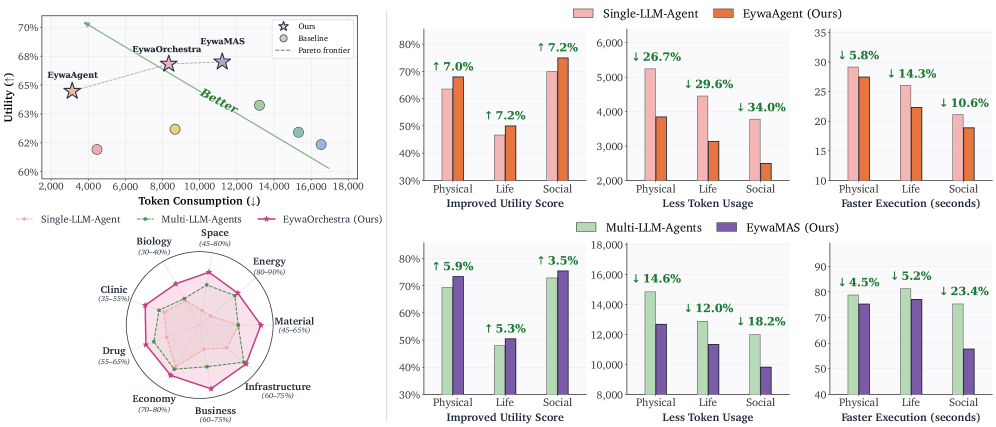
The first figure gives the paper’s practical claim: scientific agents should not force every task through language tokens. Eywa variants show higher utility with lower token consumption than language-only baselines across physical, life, and social science tasks. The caveat is that utility is aggregated across heterogeneous tasks, so the interesting story is the interface design, not one headline score.
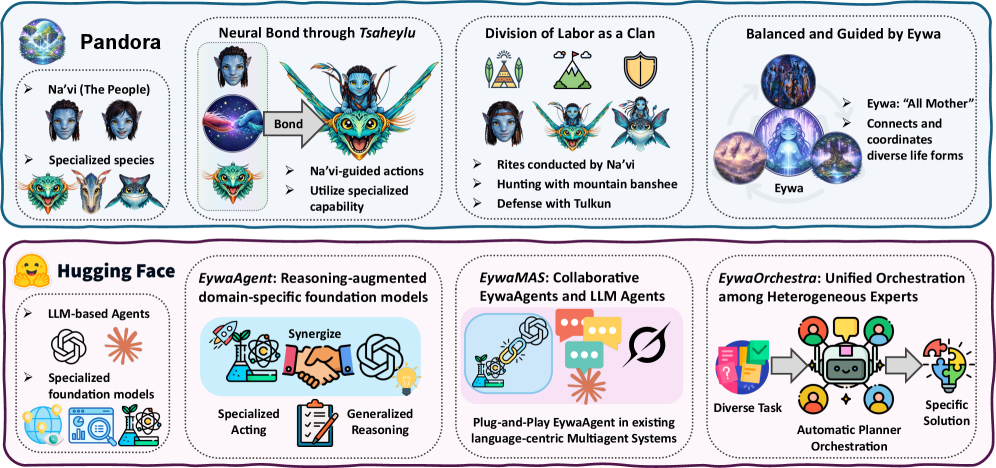
The framework diagram makes the architecture easy to understand. A language model does high-level interpretation and planning, while a domain foundation model handles specialized prediction over time series, tables, or other scientific modalities. The useful abstraction is the bidirectional interface: compile the task into a structured model call, then adapt the specialist output back into language reasoning.
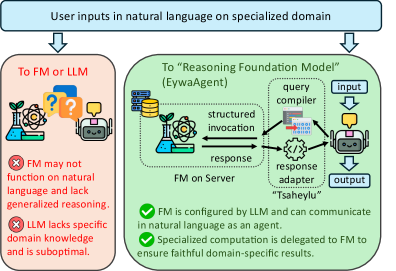
This figure zooms in on EywaAgent. The language model can skip the specialist model or invoke it through a defined input-output schema. This supports the paper’s main design claim: the agent does not need to verbalize all scientific computation, it can route the right part of the problem to a model built for that data type.
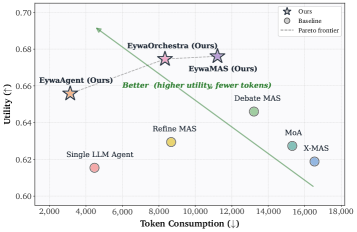
The utility-token plot is the most product-relevant evidence. Eywa tries to move the system toward better answers with fewer tokens, not just better answers with a larger reasoning trace. The caution is that the implementation uses selected foundation models and an experimental benchmark, so the exact efficiency gain should be rechecked when new domain models or noisier data sources are added.
Quick idea: Eywa gives scientific agents a structured way to collaborate with domain-specific foundation models instead of treating language as the universal interface.
Why it matters: language agents are flexible, but they are not good native interfaces for many scientific data types. A time-series foundation model, a tabular model, or a domain predictor often has the right inductive bias, while the LLM has the planning and explanation interface. Eywa’s bet is that scientific agents should compose these abilities rather than flatten everything into text.
Method walkthrough:
- Define a foundation-model and language-model interface with a query compiler and response adapter. The compiler turns task state into structured foundation-model input, and the adapter converts the output into planner-consumable context.
- Implement that interface through MCP-style servers, so each foundation model can be exposed as a service with a clear schema.
- Build EywaAgent as a coupled unit with an invoke-or-skip control policy. The agent can reason in language, call the specialist, or update its state with specialist output.
- Extend the unit to EywaMAS and EywaOrchestra, where multiple language and foundation-model agents are composed or dynamically selected by a conductor.
Evidence: the paper introduces EywaBench with physical, life, and social science domains, including material, energy, space, biology, clinic, drug, economy, business, and infrastructure subdomains. The implementation uses Chronos for time series and TabPFN for tabular prediction. Under the same backbone, EywaAgent raises overall utility from 0.6154 to 0.6558 while reducing average token use from 4,469 to 3,137. EywaMAS reaches 0.6761 overall utility, while EywaOrchestra reaches 0.6746 with lower reported latency and token use than the fixed multi-agent setup.
Why I care: this is the right direction for data agents. A capable research agent should not be proud of using more tokens to rederive what a specialist model already knows how to compute. The interface between general reasoning and specialized acting is the real design surface.
Limitations/questions: the abstraction is stronger than the current evidence. EywaBench is useful, but the selected foundation models are still a small slice of scientific modeling. I would want stress tests where the specialist model is wrong, poorly calibrated, or out of distribution, and where the language agent has to decide whether to trust the returned output.
Connection to the tracked themes: data agents, agentic scientific workflows, model context protocols, heterogeneous tool use.
AutoResearchBench: Benchmarking AI Agents on Complex Scientific Literature Discovery
Authors: Lei Xiong, Kun Luo, Ziyi Xia, Wenbo Zhang, Jin-Ge Yao, Zheng Liu, Jingying Shao, Jianlyu Chen, Hongjin Qian, Xi Yang, Qian Yu, Hao Li, Chen Yue, Xiaan Du, Yuyang Wang, Yesheng Liu, Haiyu Xu, Zhicheng Dou.
Institutions: not specified on the arXiv HTML page.
Date/Venue: April 28, 2026, arXiv preprint.
Links: arXiv | HTML | code
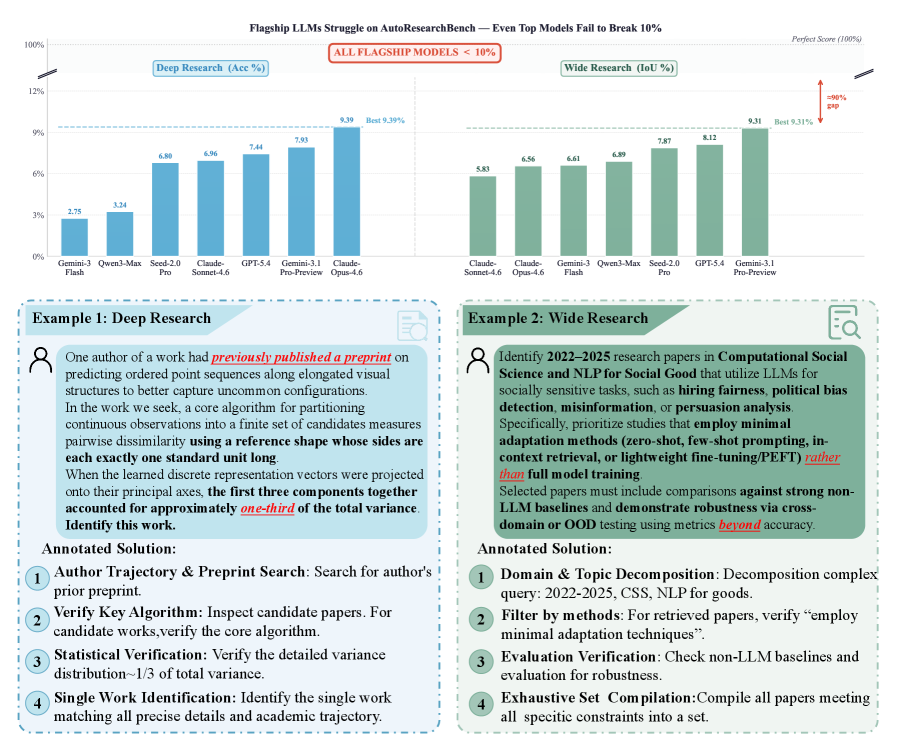

The examples figure is uncomfortable in a useful way. The benchmark is not asking an agent to find a famous paper from a title-like clue. It asks for multi-hop scientific search where decisive evidence can sit in appendix details, ablation context, citation relations, or a conjunction of weak constraints. This is much closer to how real literature work fails.
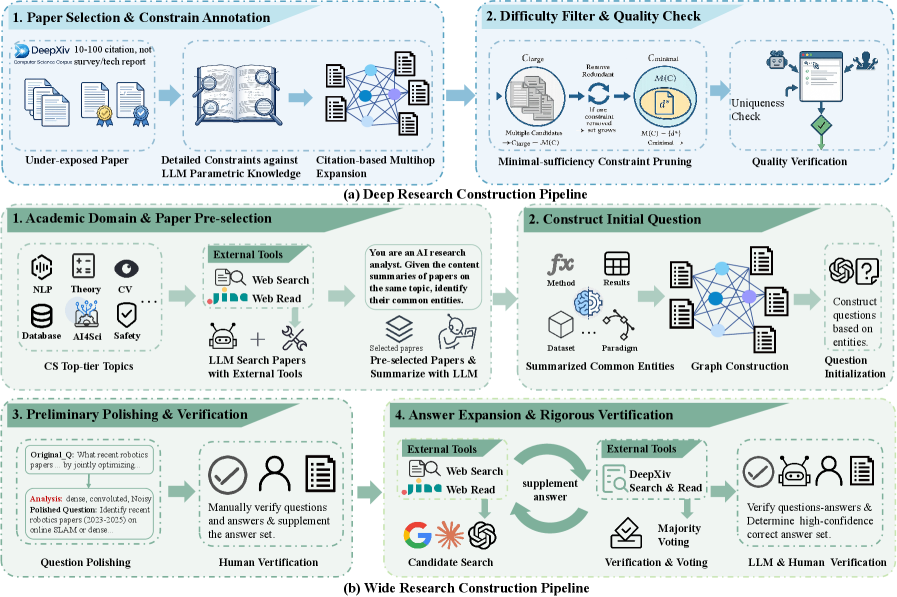
The construction pipeline explains why the benchmark is difficult. Deep Research tasks are built from full-text constraints that uniquely identify one paper or prove there is no answer; Wide Research tasks ask for an exhaustive set under strict constraints. The human verification stages matter because benchmark leakage or shallow keyword shortcuts would otherwise be easy.
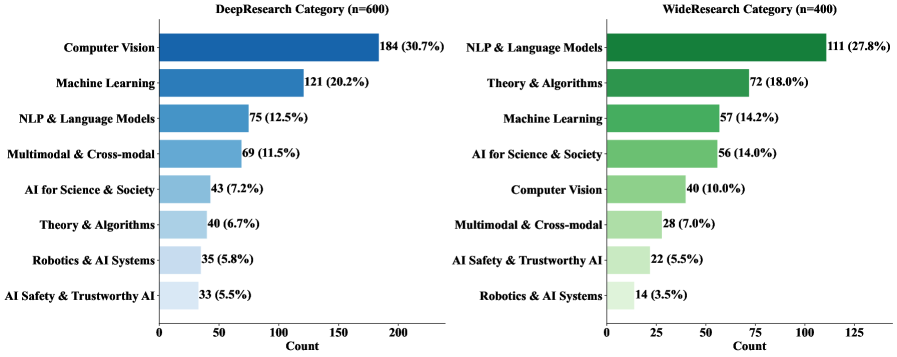
The topic distribution figure shows coverage across computer science areas rather than a single subfield. This matters because research agents can overfit to search patterns in one domain. The caveat is that the benchmark still depends on the DeepXiv corpus and its accessible full-text index, so open-web behavior may be noisier.
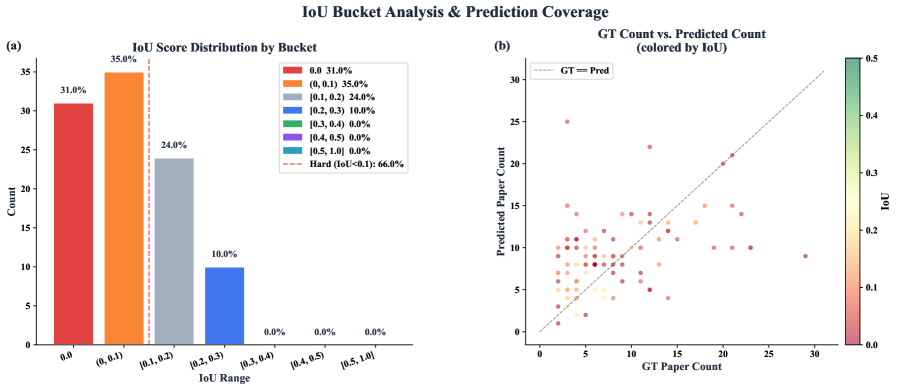
The wide-search plot makes the core failure visible: agents often retrieve something plausible but do not recover the full answer set. For a literature assistant, that is a serious bug. A partial list can look useful while silently changing the conclusion of a survey or claim-verification step.
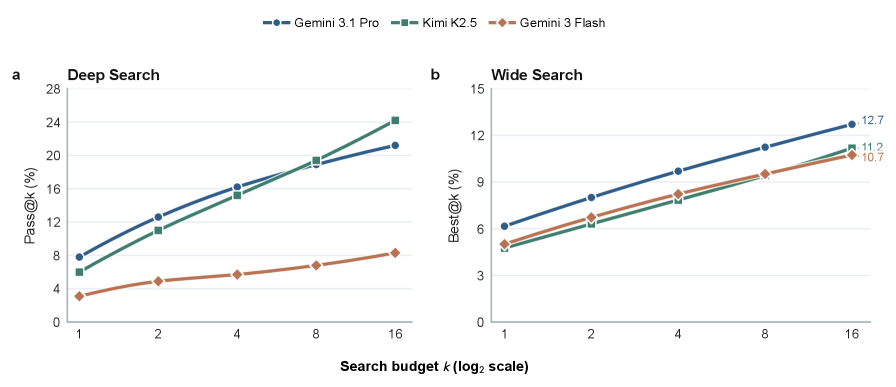
The test-time scaling figure is a warning against just giving agents more turns. The paper reports that longer trajectories do not reliably translate into better outcomes; agents can repeat similar searches, inspect plausible but wrong papers, or terminate too early. More interaction is not the same as better evidence management.
Quick idea: AutoResearchBench tests whether agents can do hard scientific literature discovery when the answer depends on full-text, multi-hop, and set-completion evidence.
Why it matters: many research-assistant demos look good because the clue is already close to the answer. Real research search is uglier. You often know a method detail, a negative result, a dataset condition, or a citation relationship, but not the target title. A useful agent must combine clues, eliminate boundary cases, and know when a search space has been exhausted.
Method walkthrough:
- Define Deep Research as exact identification of a single paper or a valid no-answer case, using tightly coupled constraints mined from full text and citations.
- Define Wide Research as constrained set completion, where the agent must retrieve all papers satisfying a scientific query without admitting out-of-scope papers.
- Build tasks through model-assisted generation plus human verification, including constraint fuzzification, pruning, adversarial search checks, and corpus-level audits.
- Evaluate ReAct-style agents using DeepXiv full-text search and compare against open-web search and end-to-end research systems.
Evidence: the benchmark contains 1,000 queries, split into 600 Deep Research and 400 Wide Research tasks, over a controlled corpus of more than three million papers. Deep Research has 90% single-answer and 10% no-answer cases; Wide Research contains 3,692 relevant papers with an average of 9.23 valid answers per query. The headline result is severe: the best reported Deep Research accuracy is 9.39% from Claude Opus 4.6, and the best Wide Research IoU is 9.31% from Gemini 3.1 Pro Preview. Open-source and proprietary models mostly remain below 10% on both tasks.
Why I care: this paper is a direct audit of tools like Paper Radar. It says that the bottleneck is not just search access. The hard part is tracking evidence through a long hypothesis space without convincing yourself too early.
Limitations/questions: the benchmark uses DeepXiv as a controlled full-text environment, which is good for reproducibility but not identical to the messy public web. I also want to see how agents behave with persistent research memory: can remembering past failed constraints help, or does it create new anchoring errors?
Connection to the tracked themes: data agents, document intelligence, autonomous research, evidence-grounded literature search.
Exploring Interaction Paradigms for LLM Agents in Scientific Visualization
Authors: Jackson Vonderhorst, Kuangshi Ai, Haichao Miao, Shusen Liu, Chaoli Wang.
Institutions: University of Notre Dame; Lawrence Livermore National Laboratory.
Date/Venue: April 30, 2026, arXiv preprint.
Links: arXiv | HTML
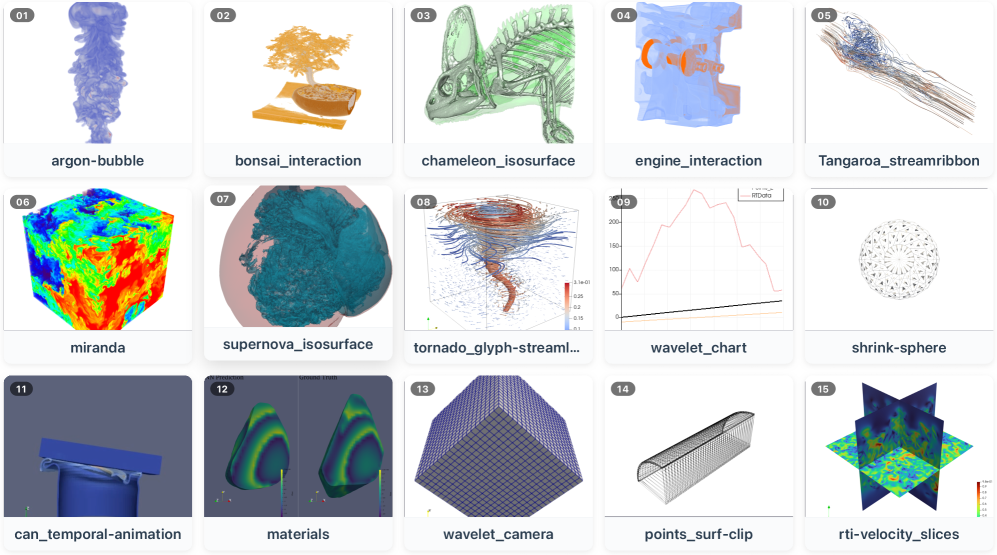

This task figure anchors the paper in actual scientific visualization work, not generic GUI clicking. The 15 ParaView tasks include loading data, applying filters, tuning parameters, setting camera views, and building multi-step visualization pipelines. The important point is that visual correctness can be semantically wrong even when the generated code runs.
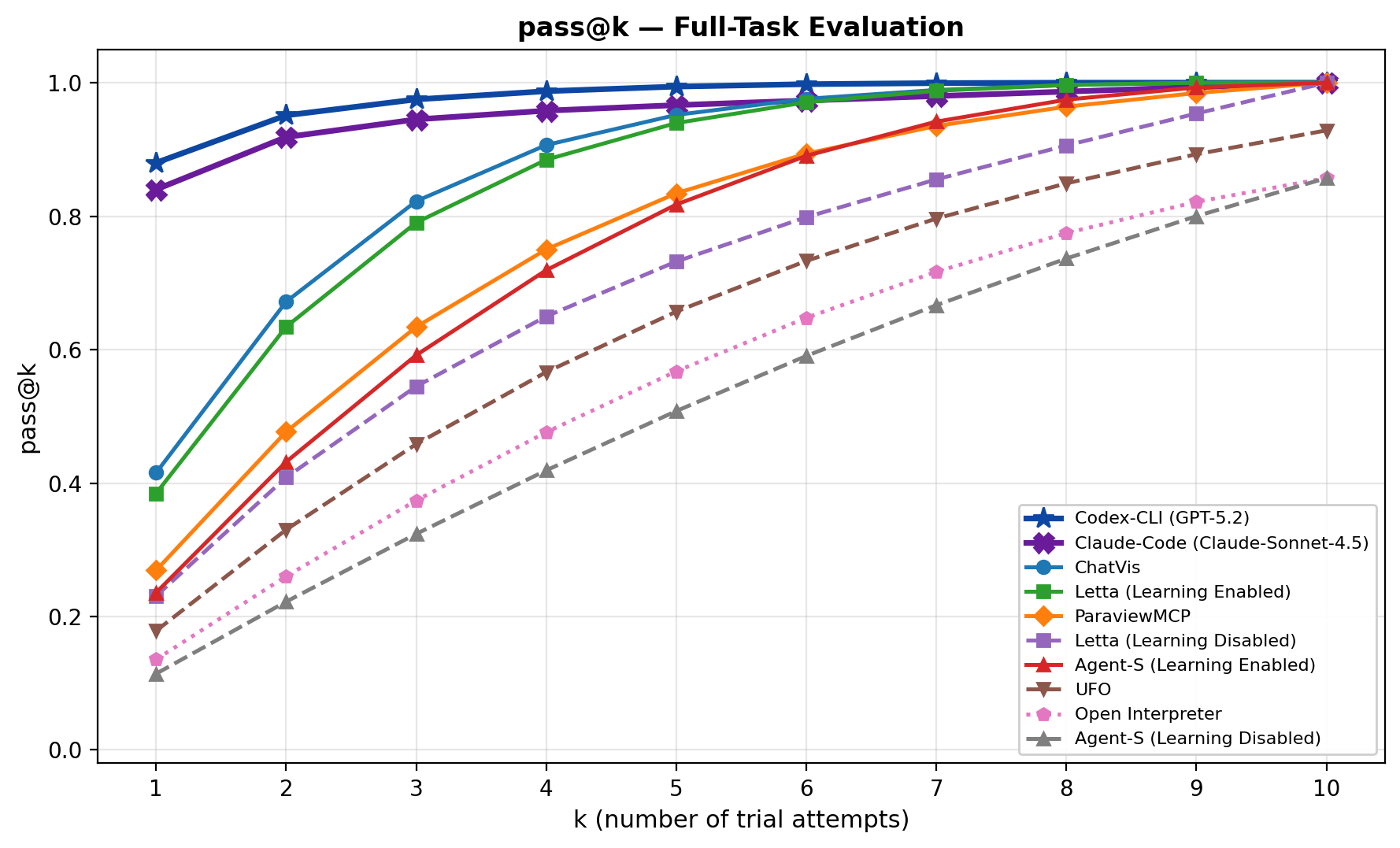
The pass-at-k curve shows that repeated attempts can make several agents look strong. Coding agents eventually approach high pass-at-k, which is not surprising when they can iterate through code and errors. The caveat is that pass-at-k is optimistic for deployed workflows unless the system has a reliable way to choose the correct run.
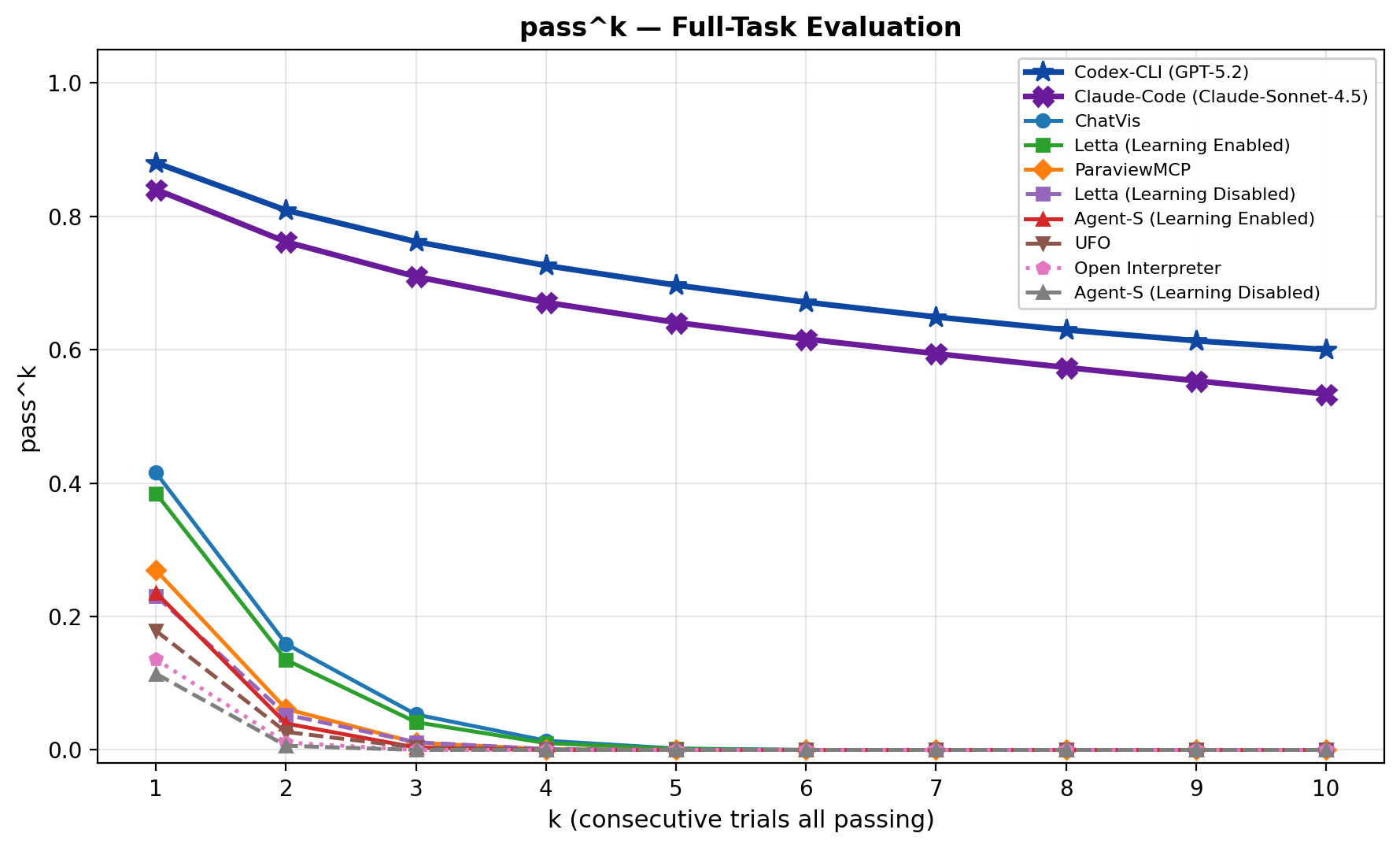
The all-run consistency curve tells the harsher story. Success is not stable across repeated trials, and consistency drops quickly. For scientific workflows, that instability matters because a user may not know which render is subtly wrong.
Quick idea: the paper compares domain-specific, GUI-based, coding-agent, and memory-enabled approaches for scientific visualization workflows, then shows that each interface fails in a different way.
Why it matters: scientific visualization is a good stress test for agents because it mixes code, visual perception, domain conventions, and intermediate state. A workflow can be executable but scientifically misleading. That makes final-answer evaluation too weak. You need to know whether the pipeline state, parameters, and rendered output match the intent.
Method walkthrough:
- Evaluate eight representative agents across three interaction paradigms: domain-specific tools such as ChatVis and ParaView-MCP, computer-use agents such as UFO and Open Interpreter, and general-purpose coding agents such as Codex CLI and Claude Code.
- Run 15 representative ParaView tasks from SciVisAgentBench, with 10 trials per task to measure both quality and variability.
- For GUI agents, add stepwise evaluation where each task is decomposed into five intermediate states, so the agent is tested on local operations without compounding prior errors.
- Compare memory-enabled and memory-disabled variants for selected agents to test whether persistent experience reduces repeated mistakes.
Evidence: coding agents are strongest on full-task completion, but expensive. The paper reports Codex CLI at 68.99 overall score and 100.0% completion, with 774.52K average input tokens per task; Claude Code reaches 66.35 and 96.0% completion, also with large token use. Domain-specific tools use far fewer tokens, for example ChatVis averages 8.17K input tokens, but their completion rates are lower. GUI agents perform poorly on full tasks, yet stepwise evaluation raises them to roughly 60-65% pass rates, suggesting that long-horizon planning is the main failure surface. Persistent memory also helps: Letta improves from 19.09 to 30.78 overall score, and Agent-S from 10.75 to 18.31.
Why I care: this paper is useful because it refuses to ask for one winner. The real takeaway is architectural: scientific agents need structured APIs, code-level flexibility, visual validation, and memory, but using any one of those alone leaves a visible hole.
Limitations/questions: the benchmark uses a small set of ParaView tasks, so I would not generalize the exact ranking too far. The more durable result is the decomposition finding. I want to see future SciVis agents expose intermediate visual states as first-class checkpoints that a human or verifier can inspect before the next step.
Connection to the tracked themes: data agents, document and visual workflow intelligence, computer-use agents, intermediate-state verification.
Reading Priority And Next Questions
My reading order after this round would be: AutoResearchBench first, because it directly audits research-assistant behavior; Intern-Atlas second, because it suggests a memory structure for scientific agents; Eywa third, because it gives a concrete interface between LLMs and scientific foundation models; and the SciVis interaction paper fourth, because it turns workflow-agent design tradeoffs into something observable.
Next questions I would track:
- Can method-evolution graphs be edited, audited, and versioned by domain experts rather than frozen after LLM extraction?
- Can scientific agents learn when a specialist foundation model is out of distribution or miscalibrated?
- Can literature-search agents keep a durable hypothesis ledger so failed constraints help later search instead of wasting turns?
- Can workflow agents expose intermediate scientific states, not only final files, as the main object of verification?